Panels currently available
Single stranded RNA ligases
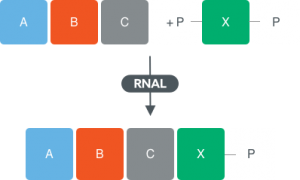
- Diverse panel of enzymes that contain thermophilic homologues that work up to 65-70 °C
- Substrate preference single stranded RNA
- Enzymes can accept unnatural 2’ modifications of 3’,5’-diphosphate donors
- Single nucleoside addition
- Coupling of oligomers
- Panel of available 96 enzymes
Double stranded RNA ligases
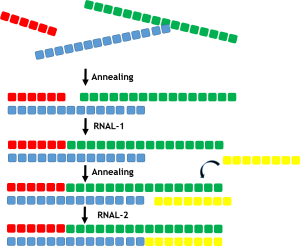

- Diverse panel of ligases from bacterial and viral sources
- Complementary base-pairing allows for blockmers to anneal to form duplex followed by ligation of adjacent blockmers by dsRNAL
- Panel of available 96 enzymes
Alkaline phosphatases
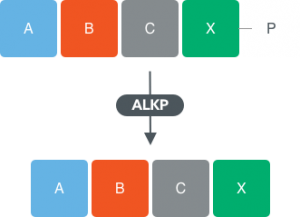

- For the removal of terminal phosphate to allow further extension of oligonucleotide
- Highly thermostable and can be heat purified
- Panel of available 96 enzymes
Polynucleotide phosphorylases
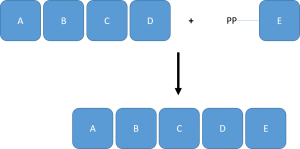

- PNPase catalyzes the reversible polymerization of ribonucleoside diphosphates to polyribonucleotide with the release of inorganic phosphate
- Panel of available 96 enzymes
Nucleoside deoxyribosyltransferases
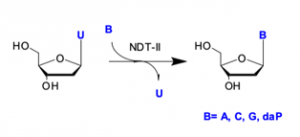

- Catalyses the cleavage of the glycosidic bond of 2′-deoxyribonucleosides and the transfer of the deoxyribosyl moiety to an acceptor purine or pyrimidine base
- Useful for synthesis of modified nucleoside building blocks.
- Panel of available 50 enzymes
Phosphorylases
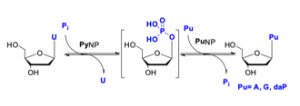

- Phosphorylases (PPL) are known to be more promiscuous than NDTs in 2’ position and are known to be active against nucleosides containing F, NH2, OH and H in 2’ position.
- Panel of 25 purine and 25 pyrimidine nucleotide phosphorylases
Polynucleotide kinases
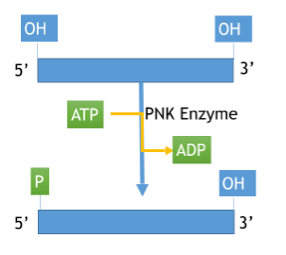

Polynucleotide kinase (PNK)
- Transfers phosphate from ATP to 5’OH of RNA blockmers.
- Panel of 50 enzymes available
Panels in development
- DNA ligase
- Phosphodiesterase
- Phosphotriesterase
- Terminal deoxynucleotidyl transferase
- DNA polymerases
- Phosphoramidases


